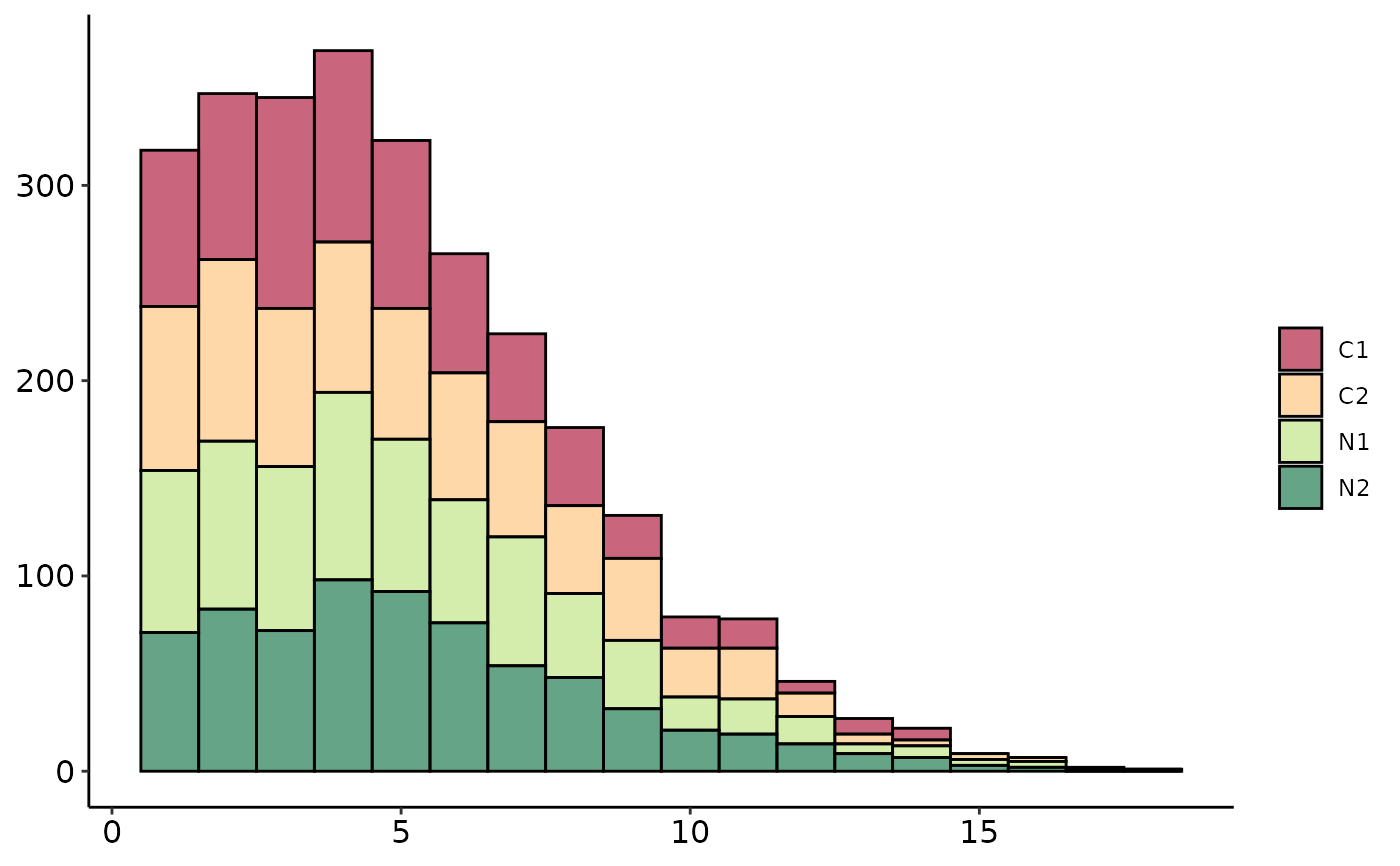
Coverage QC Plots
plot_coverage.RdCoverage QC Plots
plot_coverage( m, type = c("hist", "dens"), pheno = NULL, perGroup = FALSE, lim = 100, size.lim = 1e+06, col_palette = "RdYlGn" )
Arguments
| m | Input |
|---|---|
| type | Choose between 'hist' (histogram) or 'dens' (density plot). |
| pheno | Column name of colData(m). Will be used as a factor to color different groups in the plot. |
| perGroup | Color the plots in a sample-wise manner? |
| lim | Maximum coverage value to be plotted. |
| size.lim | The maximum number of observarions (sites*samples) to use. If the dataset is larger that this, random sites will be selected from the genome. |
| col_palette | Name of the RColorBrewer palette to use for plotting. |
Value
ggplot2 object